 Benjamin Balluff, Sandra Rauser, Stephan Meding, Mareike Elsner, Cedrik Schöne, Annette Feuchtinger, Christoph Schuhmacher, Alexander Novotny, Uta Jütting, Giuseppina Maccarrone, Hakan Sarioglu, Marius Ueffing, Herbert Braselmann, Horst Zitzelsberger, Roland M. Schmid, Heinz Höfler, Matthias P. Ebert and Axel Walch
Benjamin Balluff, Sandra Rauser, Stephan Meding, Mareike Elsner, Cedrik Schöne, Annette Feuchtinger, Christoph Schuhmacher, Alexander Novotny, Uta Jütting, Giuseppina Maccarrone, Hakan Sarioglu, Marius Ueffing, Herbert Braselmann, Horst Zitzelsberger, Roland M. Schmid, Heinz Höfler, Matthias P. Ebert and Axel Walch
Am J Pathol. (2011), 179(6), 2720-2729
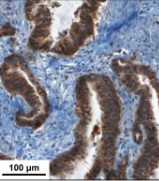
Proteomics-based approaches allow us to investigate the biology of cancer beyond genomic initiatives. We used histology-based matrix-assisted laser desorption/ionization (MALDI) imaging mass spectrometry to identify proteins that predict disease outcome in gastric cancer after surgical resection. A total of 181 intestinal-type primary resected gastric cancer tissues from two independent patient cohorts were analyzed. Protein profiles of the discovery cohort (n = 63) were directly obtained from tumor tissue sections by MALDI imaging. A seven-protein signature was associated with an unfavorable overall survival independent of major clinical covariates. The prognostic significance of three individual proteins identified (CRIP1, HNP-1, and S100-A6) was validated immunohistochemically on tissue microarrays of an independent validation cohort (n = 118). Whereas HNP-1 and S100-A6 were found to further subdivide early-stage (Union Internationale Contre le Cancer [UICC]-I) and late-stage (UICC II and III) cancer patients into different prognostic groups, CRIP1, a protein previously unknown in gastric cancer, was confirmed as a novel and independent prognostic factor for all patients in the validation cohort. The protein pattern described here serves as a new independent indicator of patient survival complementing the previously known clinical parameters in terms of prognostic relevance. These results show that this tissue-based proteomic approach may provide clinically relevant information that might be beneficial in improving risk stratification for gastric cancer patients.
No responses yet